Once the molecule file is fully loaded, the image at right will become live. At that time the "activate 3-D" icon ![]() will disappear.
will disappear.
Bromoflouride.
Two different geometry optimizations were used to calculate geometries using the highest three theories of optimization. PM3 was the optimization used for all the other three theories.
AM1 was the other geometry optimization calculation but it was not used for any of the other theories used.
Once the PM3 geometry was found the 3-21G theory was used to calculate the optimized geometry. It was the lowest theory for geometry optimization.
6-31G was the second level of geometry optimization theory. This theory gave a geometry optimization closest to the theoretical value for bond length which was found to be 0.1759nm
DZV was the highest level of theory used to determine geometry optimization but didn't give a value as close to the theoretical bond length as the 6-31G.
The highest occupied molecular orbital was at orbital 22. That number was calculated by summing all the electrons in both molecules dividing the number of electrons by 2.
The lowest unoccupied molecular orbital was at orbital 23 which was calculated by adding one more electron to the HOMO. This would be the next orbital to be occupied when an electron from the HOMO was excited enough to increase to the next orbital.
The electrostatic potential is shown here, where the blue represents the highest potential and the red shows the lowest potential. The other colors represent the intermediate potentials the more blue the higher the potential.
The partial atomic charges are shown in black, and created by an asymmetric distribution of electrons in the chemical bond.
Table 1: Valence Shell Orbitals on the bromoflouride atom. The S-orbital is the lowest energy increasing to the P-orbital and shows the bonding and anti-bonding characteristics.
| Types of Bonding |
Orbital |
| S sigma bonding |
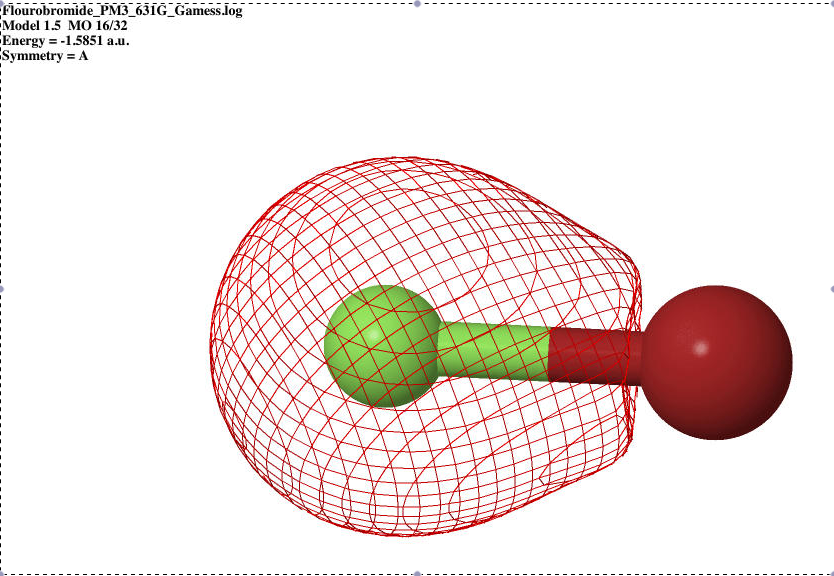 |
| S sigma anti-bonding |
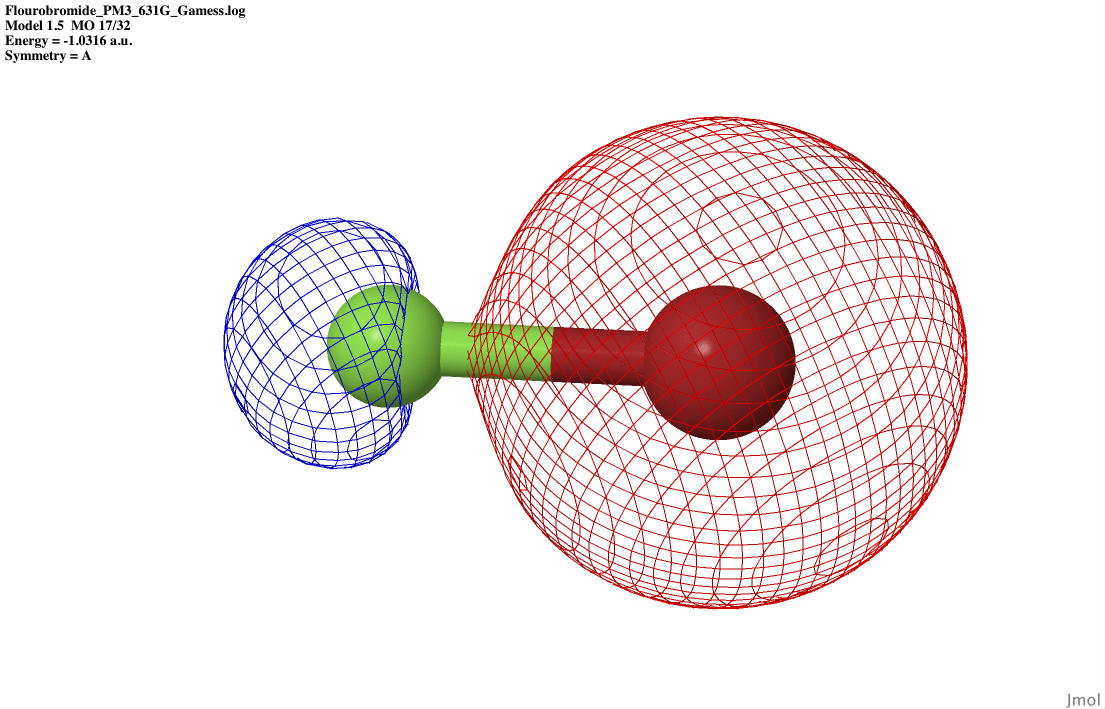 |
| P bonding |
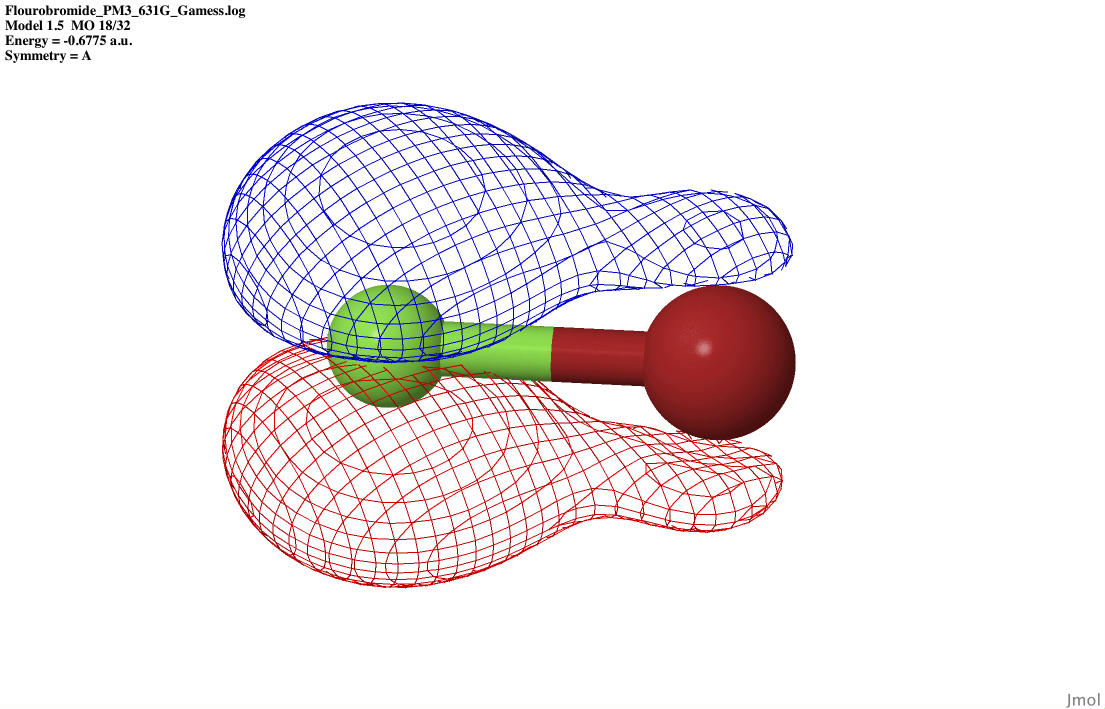 |
| P anti-bonding |
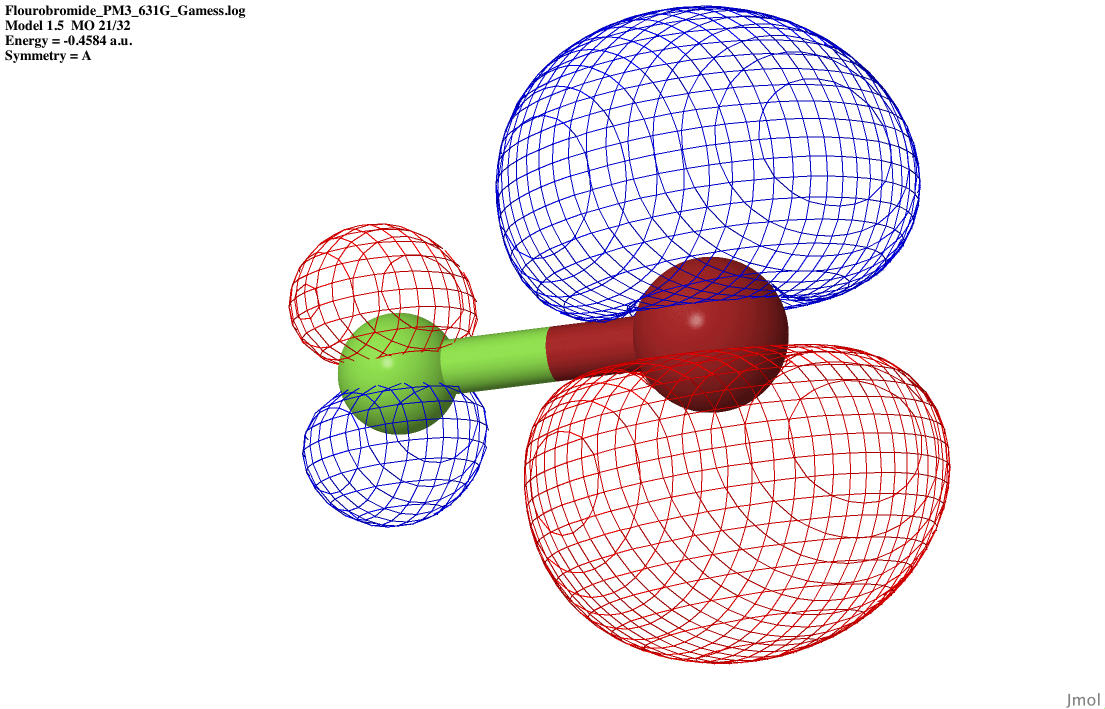 |
| P2 bonding |
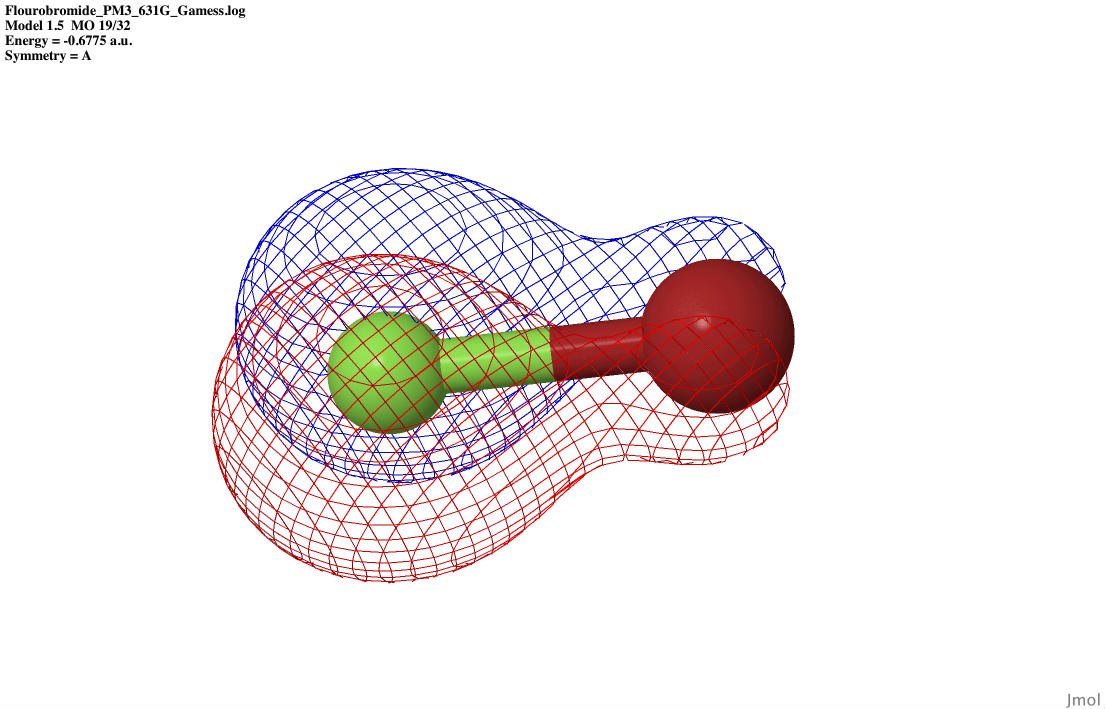 |
| P2 anti-bonding |
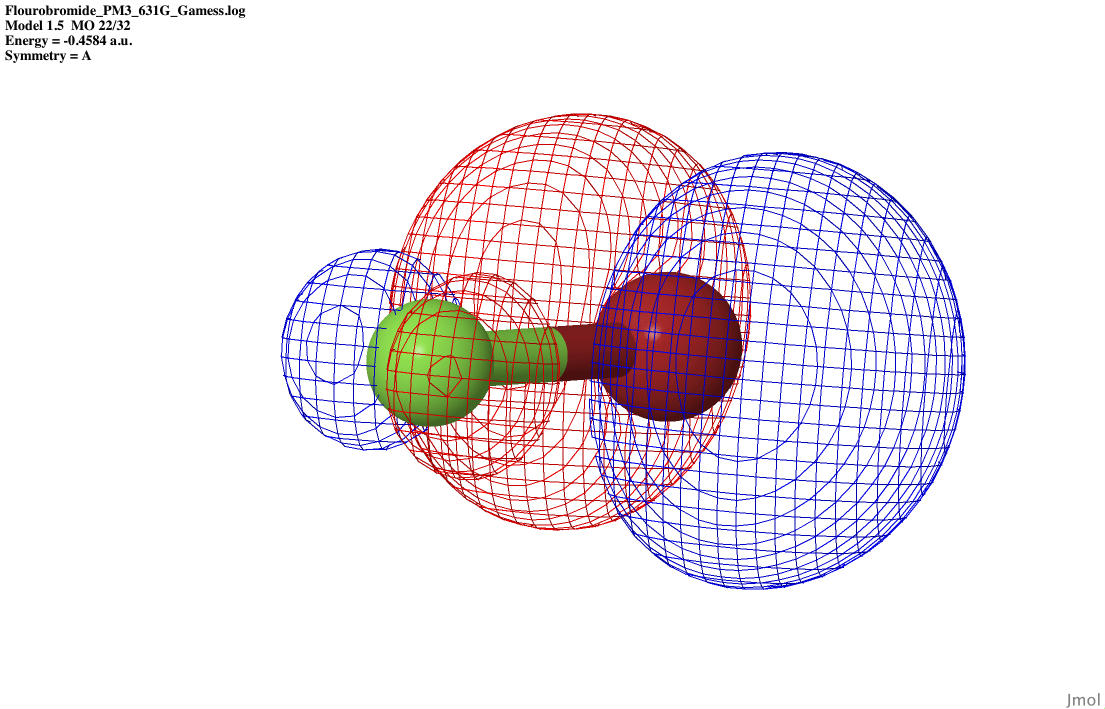 |
The potential energies during bond stretching were different depending on the level of theory that was used. The lower the potential energy the more accurate the level of theory was in predicting the lowest potential energy. The "bumps" on the graph shows the interactions of the electrons that the theory doesn't take into account in the calculations. If the potential energy was measured experimentally the "bump" would not appear on the graph.
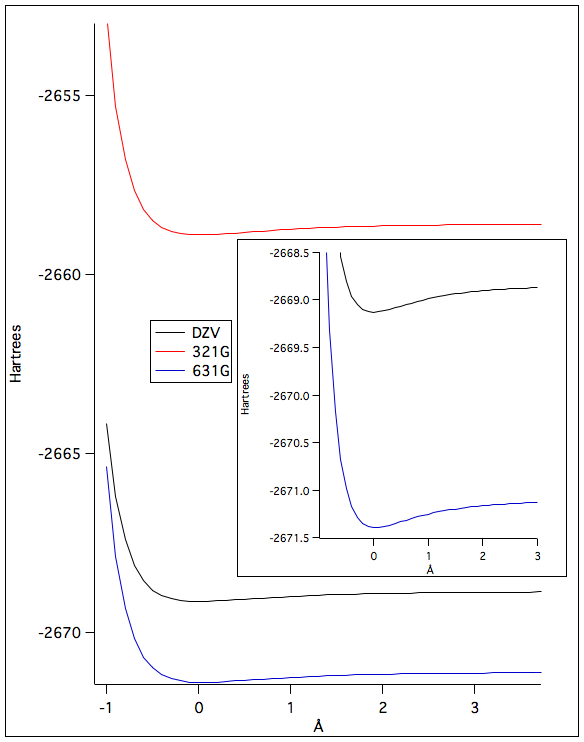
Figure 1: The potential energies at different bond lengths calculated using different levels of theory. The "bumps" for the 6-31G and DZV theory were small so a zoomed in graph was also added to show the dip in the graphs. The two graphs were manufactured on IGOR Pro.
The vibrational frequency using the 6-31G theory was found to be 691cm^-1. Using NIST the theoretical vibrational frequency was found to be 720 cm^-1 which gave an error of 4.20%.
Using all the levels of theory the dipole moments were found and compared to the theoretical dipole moment that was found on NIST.
Table 2: Dipole moments found using each of the levels of theory. Percent error for each of the measurements were calculated comparing experimental measurements to the theoretical found on NIST.
| Theory |
Dipole Moments (D) |
Error in Experimental Calculations (%) |
| 3-21G |
1.768 |
10.6 |
| 6-31G |
1.730 |
13.0 |
| DZV |
1.809 |
8.07 |
From the data it can be seen that the DZV theory gave the best dipole moment value. Since DZV is the highest theory that was used it is not surprising that it gave the dipole closest to the theoretical. The 6-31G theory should have been closer to the theoretical than the 3-21G theory since it uses more basis sets in the calculations, the more basis sets a theory uses the more accurate the values it calculates are.
You may look at any of these intermediate views again by clicking on the appropriate button.
Page skeleton and JavaScript generated by export to web function using Jmol 14.2.12_2015.01.22 2015-01-22 21:48 on Mar 2, 2015.
This will be the viewer

If your browser/OS combination is Java capable, you will get snappier performance if you use Java