Once the molecule file is fully
loaded, the image at right will become live. At that time the
"activate 3-D" icon ![]() will disappear.
will disappear.
Monofluoroamine (NH2F)
The geometry optimizations for the three
highest levels of theory are shown below. The following
views show both the bond lengths and angles.
6-21G was the lowest level of theory used
for geometry optimization.
DZV was the highest
level of theory used for geometry optimization.
In Table 1, the bond
lengths can be compared to their literature values.
Table 2 can be used to
compare the bond angles.5
Table 1: Bond lengths from NIST database.
|
Bond Type |
Bond
Length (angstroms) |
|
N-H |
1.0085 |
|
N-F |
1.3828 |
Table 2: Bond angles from NIST database.
|
Bond Type |
Bond Angle (degrees) |
|
F-N-H |
103.401 |
|
H-N-H |
105.393 |
|
F-N-H |
103.401 |
The partial atomic charge on each atom is
shown here. They are created by the asymmetric distribution
of electrons in a chemical bond.
The vibrational frequencies were calculated using the highest level of theory. These frequencies would be the peaks on an infrared spectroscopy spectrum of monofluoroamine. Table 3 lists these frequencies in comparison with literature values.6
Table 3: Tabulated values for the frequencies and types of modes of each kind.
| Type of mode |
DZV Frequency (cm-1) |
Literature Frequency (cm-1)
|
| NF stretch |
1015.86 |
934 |
| NH2 wag |
1255.06 |
1244 |
| NH2 scissor |
1787.07 |
1568 |
| NH2 symmetric stretch |
3730.38 |
3269 |
| NH2 asymmetric stretch |
3876.36 |
3346 |
The following buttons show the calculated vibrational frequencies using DZV. Clicking them will show each type of vibration for the molecule.
The dipole moment was calculated at five
different levels of theory to determine the best value. The
best theory was DZV with a diffusion of 100 which gave 2.610
debyes. When compared the a literature value of 2.615
debyes, the error was 0.191%.7 Table 4 lists the
results from each theory.
Table 4: Tabulated values for the dipole moment in monofluoroamine. Note that the diffusion was a variation on #D heavy atom polarization functions, #F heavy atom polarization functions, and # light atom polarization functions.
Table 4: Tabulated values for the dipole moment in monofluoroamine. Note that the diffusion was a variation on #D heavy atom polarization functions, #F heavy atom polarization functions, and # light atom polarization functions.
| 6-21G |
6-31G |
PM3 |
AM1 |
DZV |
Diffusion (#D, #F, # light) |
| 2.837 |
3.043 |
1.916 |
2.138 |
3.105 |
No diffusion |
| 2.846 |
2.549 |
2.610 |
100 |
||
| Error |
2.516 |
2.580 |
110 |
||
| 2.776 |
2.534 |
2.589 |
101 |
You may look at any of these intermediate views again by clicking on the appropriate button.
Page skeleton
and JavaScript generated by export to web function using Jmol 14.2.15_2015.07.09
2015-07-09 22:22 on Mar 6, 2016.
This will be the viewer 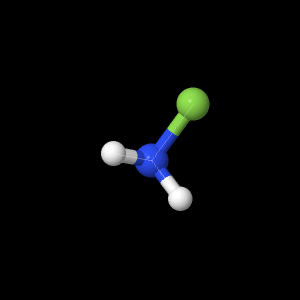

If your
browser/OS combination is Java capable, you will get snappier
performance if you use Java